One of the keys to commercialization of advanced biofuels is the development of cost-competitive ways to extract fermentable sugars from lignocellulosic biomass. The use of enzymes from thermophiles – microbes that thrive at extremely high temperatures and alkaline conditions – holds promise for achieving this. Finding the most effective of these microbial enzymes, however, has been a challenge. That challenge has now been met by a collaboration led by researchers with the U.S. Department of Energy (DOE)’s Joint BioEnergy Institute (JBEI).
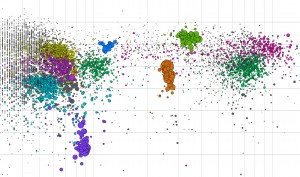 |
| The microbial world of biomass deconstruction became more clear with a JBEI/JGI/EMSL study of a thermophillic bacterial consortium adapted to switchgrass. This splatterplot is a visual representation of the consortium’s metagenome. (Image courtesy of Patrik D’haeseleer, JBEI) |
Working with a compost-derived consortium of thermophillic bacterium adapted to grow on switchgrass, a leading potential fuel crop, and using a combination of metagenomic and metaproteomic technologies, the collaboration has identified individual microbial species whose enzymes were the most active in deconstructing the switchgrass biomass. Major institutes in addition to JBEI participating in this collaboration included DOE’s Joint Genome Institute (JGI), and EMSL, the Environmental Molecular Sciences Laboratory, a national scientific user facility at Pacific Northwest National Laboratory (PNNL).
“This marks the first time that the functional roles of individual microbial populations within a consortium have been linked with specific enzyme activities, in this case cellulase and hemicellulase,” says Steven Singer, director of JBEI’s microbial communities program. “Since these activities are broadly relevant to biofuel production, this is one of the first real-world applications being met by combining metagenomics and metaproteomics.”
Singer, who is also a research scientist with Lawrence Berkeley National Laboratory (Berkeley Lab)’s Earth Sciences Division, is the senior author of a paper describing this research in the journal PLOS One titled “Proteogenomic Analysis of a Thermophilic Bacterial Consortium Adapted to Deconstruct Switchgrass.” Co-authors are Patrik D’haeseleer, John Gladden, Martin Allgaier, Patrik Chain, Susannah Tringe, Stephanie Malfatti, Joshua Aldrich, Carrie Nicora, Errol Robinson, Ljiljana Paša-Tolić, Philip Hugenholtz and Blake Simmons.
Advanced biofuels – liquid transportation fuels synthesized from the sugars in cellulosic biomass – offer a clean, green and renewable alternative to gasoline, diesel and jet fuels. However, unlike the simple sugars in corn grain, the cellulose and hemicellulose in biomass are difficult to extract in part because they are embedded in a tough woody material called lignin. Thermophilic microbes are believed to be a rich source of cellulase and hemicellulase enzymes for lignocellulosic biomass deconstruction that are active at elevated temperatures and in the presence of pretreatment chemicals such as ionic liquids.
“Natural microbial communities that deconstruct biomass, such as those found in cow rumen or compost piles, are often too complex to decipher roles for individual microbial populations,” says Singer. “However, enrichment cultures established with defined substrates and at constant temperatures offer the possibility of simplifying these complex microbial communities and identifying functional roles for specific populations within the community.”
As part of their efforts to develop a cost-effective way to deconstruct lignocellulosic biomass into sugars for fuel, researchers in JBEI’s Deconstruction Division cultivated the switchgrass-feeding, compost-derived consortium of thermophiles.
“Using pretreated switchgrass at temperatures up to 80 degrees Celsius, we demonstrated that this consortium is an excellent source of enzymes for the development of enzymatic cocktails tailored to biorefinery processing conditions,” says Blake Simmons, a chemical engineer who heads JBEI’s Deconstruction Division and was a member of this research collaboration.
To identify the functional roles of community members within the switchgrass-feeding consortium, Singer, Simmons and their colleagues first used shotgun sequencing, a powerful metagenomics technique that enabled them to determine the metabolic potential of all the members of the consortium. They then used metaproteomic measurements to identify those enzymes, predicted by metagenomic analysis, that were actually produced by the microbial community.
 |
| Steve Singer directs the Joint BioEnergy Insitute (JBEI)’s microbial communities program. (Photo by Roy Kaltschmidt, Berkeley Lab) |
“Doing metagenomics by shotgun sequencing is a bit like raiding a toy store and tossing hundreds of jigsaw puzzles onto a pile,” says Patrik D’haeseleer, a computational systems biologist who holds appointments with both JBEI and the Lawrence Livermore National Laboratory and is lead author of the PLOS One paper.
“Each individual puzzle piece may carry some useful information, but you only start to see the bigger picture once you reassemble the pieces,” D’haeseleer says. “Our collaborators at JGI used deep sequencing of millions of small pieces of DNA, and generated a partial assembly based on direct matches between the pieces. We developed a novel phylogenetic binning method to separate those partially assembled pieces into the major bacterial genomes in the consortium. This allowed us to then model the metabolic potential of all those members of the consortium.”
Analysis of metagenomic sequencing data identified the most abundant microbial populations in the consortium to be closely related strains of Thermus thermophilus and Rhodothermus marinus. However, based on the assigned fractions of the switchgrass deconstruction proteome, the strains showing the most active role in switchgrass deconstruction were Gemmatimonadetes and Paenibacillus. By comparison, the more numerous Rhodothermus strain contributed fewer enzymes to biomass deconstruction.
“By leveraging the unique capabilities of the JGI and EMSL with those at JBEI, we’re developing a more comprehensive functional understanding of how microbial consortia work to breakdown lignocellulose, and identifying the genes and enzymes that are responsible for this deconstruction,” Simmons says. “The list of genes and enzymes generated by this study has been placed into our expression pipeline and are being used to develop optimized cocktails that are capable of generating high sugar yields from pretreated lignocellulosic biomass.”
This research was supported by the DOE Office of Science.
About JBEI:
JBEI is one of three Bioenergy Research Centers established by the DOE’s Office of Science in 2007. It is a scientific partnership led by Berkeley Lab and includes the Sandia National Laboratories, the University of California campuses of Berkeley and Davis, the Carnegie Institution for Science, and the Lawrence Livermore National Laboratory. DOE’s Bioenergy Research Centers support multidisciplinary, multi-institutional research teams pursuing the fundamental scientific breakthroughs needed to make production of cellulosic biofuels, or biofuels from nonfood plant fiber, cost-effective on a national scale.
DOE’s Office of Science:
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the Unites States, and is working to address some of the most pressing challenges of our time.
Source
Lawrence Berkeley National Laboratory, press release, 2013-07-25.
Supplier
Environmental Molecular Sciences Laboratory (EMSL)
Joint BioEnergy Institute (JBEI)
Lawrence Berkeley National Laboratory
Pacific Northwest National Laboratory PNNL
US Department of Energy (DoE)
US Department of Energy Joint Genome Institute (DOE JGI)
US DOE Office of Science (SC)
Share
Renewable Carbon News – Daily Newsletter
Subscribe to our daily email newsletter – the world's leading newsletter on renewable materials and chemicals










